Modern medicine often fails people with rare disorders, leaving patients to navigate years of uncertainty before a definitive diagnosis. For families and clinicians alike, earlier answers can change the entire trajectory of care, enabling timely treatment and realistic planning.
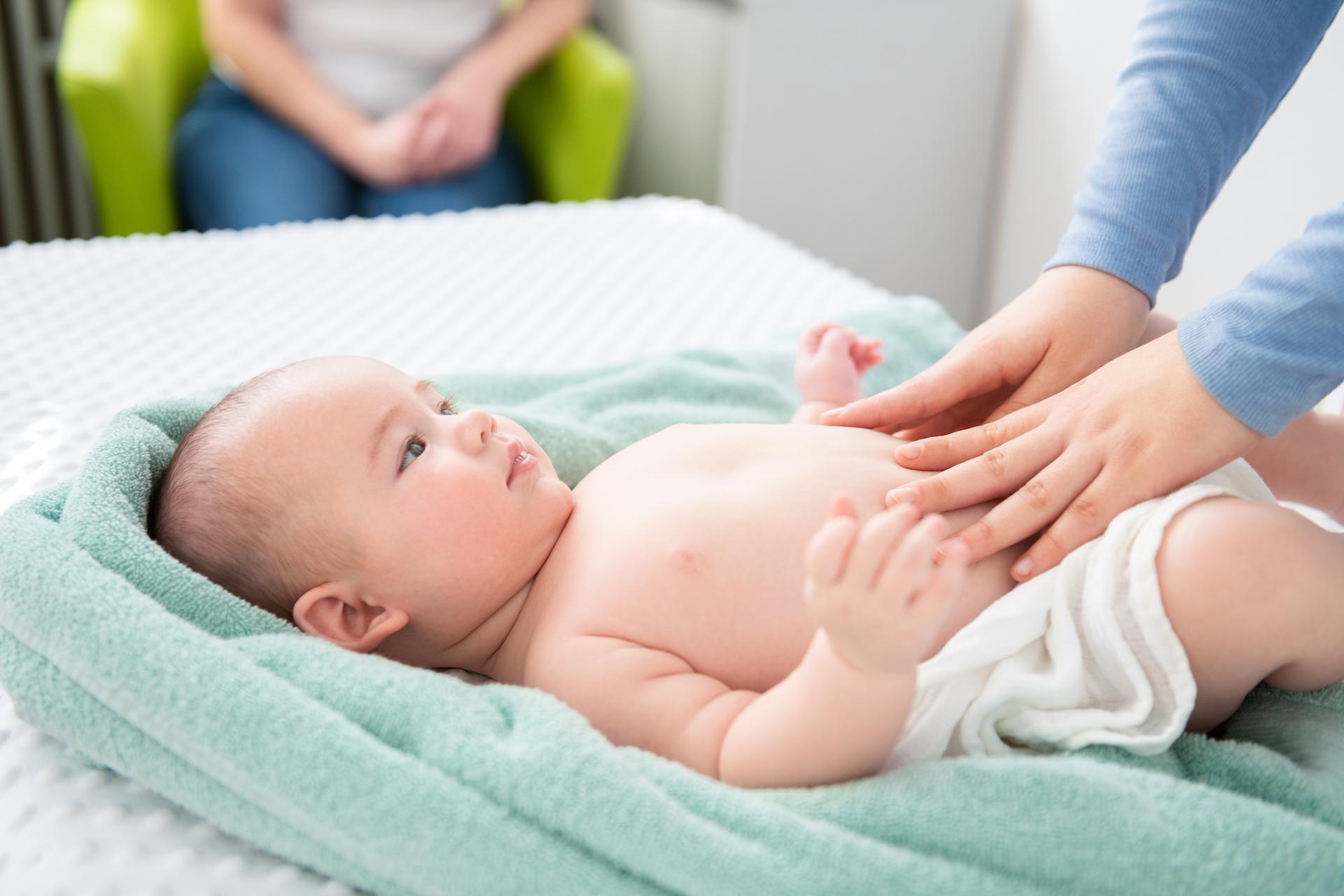
A minimally invasive blood test opens a new diagnostic path for rare genetic diseases. © busracavus, iStock
From DNA to proteins: a shift in focus
Most rare genetic diseases are investigated through whole‑genome or exome sequencing, which delivers answers in only 30–50% of cases. A new Australian approach flips the script by reading the body’s protein output, rather than the DNA blueprint itself. By interrogating proteins, researchers can see how sequence changes alter cellular function, providing clues that pure DNA data can miss.
The method profiles more than 8,000 proteins in peripheral blood cells, capturing signals from over half of known Mendelian and mitochondrial disease genes. With just one milliliter of blood, results are returned in under three days. That speed and minimal invasiveness make the tool promising for newborns, complex adult cases, and longitudinal monitoring across a lifetime.
As lead author Daniella Hock notes, “Our platform can screen thousands of proteins and potentially identify both known and previously unrecognized disease drivers.” By connecting molecular changes to protein networks, the assay helps convert genetic variants of uncertain significance into interpretable findings.
Why this could transform care
Faster, clearer answers can shorten the diagnostic odyssey, reduce repeated testing, and unlock earlier interventions. Because proteomic readouts reflect what cells are actually doing, they can illuminate disease pathways even when DNA results are ambiguous.
Key advantages include:
- Earlier, minimally invasive diagnosis, enabling timely specialty referral
- Functional context for cryptic genetic variants, improving clinical confidence
- Broad coverage across thousands of rare conditions, not just single‑disease panels
- Turnaround measured in days, not weeks or months
- Potential use in neonatal, prenatal, or preimplantation settings, with appropriate counseling
For families facing progressive or life‑limiting conditions, weeks can matter for supportive therapies. In metabolic or mitochondrial disorders, earlier identification can prevent irreversible damage, improve growth, and reduce avoidable hospitalizations.
The economics of earlier answers
Rare diseases collectively affect an estimated 300 million people worldwide, spanning more than 7,000 distinct entities. The cumulative cost of delayed diagnosis—specialist visits, unnecessary procedures, and misdirected therapies—is enormous for both families and health systems. Researchers suggest the new assay could be priced on par with existing mitochondrial disease tests, yet it potentially addresses thousands more conditions.
Economies arise not only from broader utility, but from replacing serial testing with a single, information‑rich screen. By guiding targeted confirmatory workups, the approach can minimize redundant panels and reduce inpatient diagnostic days. Health services gain when answers arrive sooner and pathways become predictable, making care more efficient and equitable.
Where the evidence stands
The platform is currently undergoing clinical evaluation, including a 300‑participant cohort with diverse genetic disorders. Investigators aim to measure sensitivity, specificity, and real‑world usability across different ages, phenotypes, and comorbid states. Reproducibility across laboratories and instruments is a crucial benchmark, as is performance in mixed or complex presentations.
If validated, the test will still need clear clinical guidelines, integration into electronic records, and payer policies that reflect its value. Multidisciplinary teams—genetics, neurology, metabolic medicine, and primary care—will be essential for translating results into tailored management.
Challenges to solve before scale
Proteomes are dynamic, shaped by time, tissue, and environment, so pre‑analytical variables must be rigorously standardized. Signal interpretation requires high‑quality reference databases, careful control of confounders, and transparent reporting of uncertainty to prevent over‑ or under‑diagnosis. Ethical safeguards around prenatal or preimplantation use demand robust consent and clear communication.
Equity is another imperative: access should extend beyond academic centers, and validation must include diverse genetic backgrounds. Finally, regulatory clearance will hinge on demonstrated clinical utility, robust analytical performance, and a repeatable end‑to‑end workflow.
A pragmatic path to precision
What makes this advance compelling is its practical blend of speed, breadth, and function. By moving from static DNA code to dynamic protein readouts, clinicians gain a bridge between genotype and phenotype. Even where it does not deliver a definitive label, it can narrow the differential, prioritize confirmatory tests, and point toward plausible therapies.
No single tool will end the diagnostic odyssey, but this one could make it significantly shorter. If ongoing trials confirm accuracy, and if implementation is matched by ethical, economic, and access‑focused policies, the result could be a tangible shift from reactive to predictive, from fragmented to precise. For millions living in the shadows of undiagnosed disease, that is a meaningful and timely step.